AI drug discovery models fail due to a fundamental lack of high-quality, labeled biological data. Synthetic data generation is the only scalable solution to train robust target identification models.
Blog
How Synthetic Data Trains Robust Target Identification Models
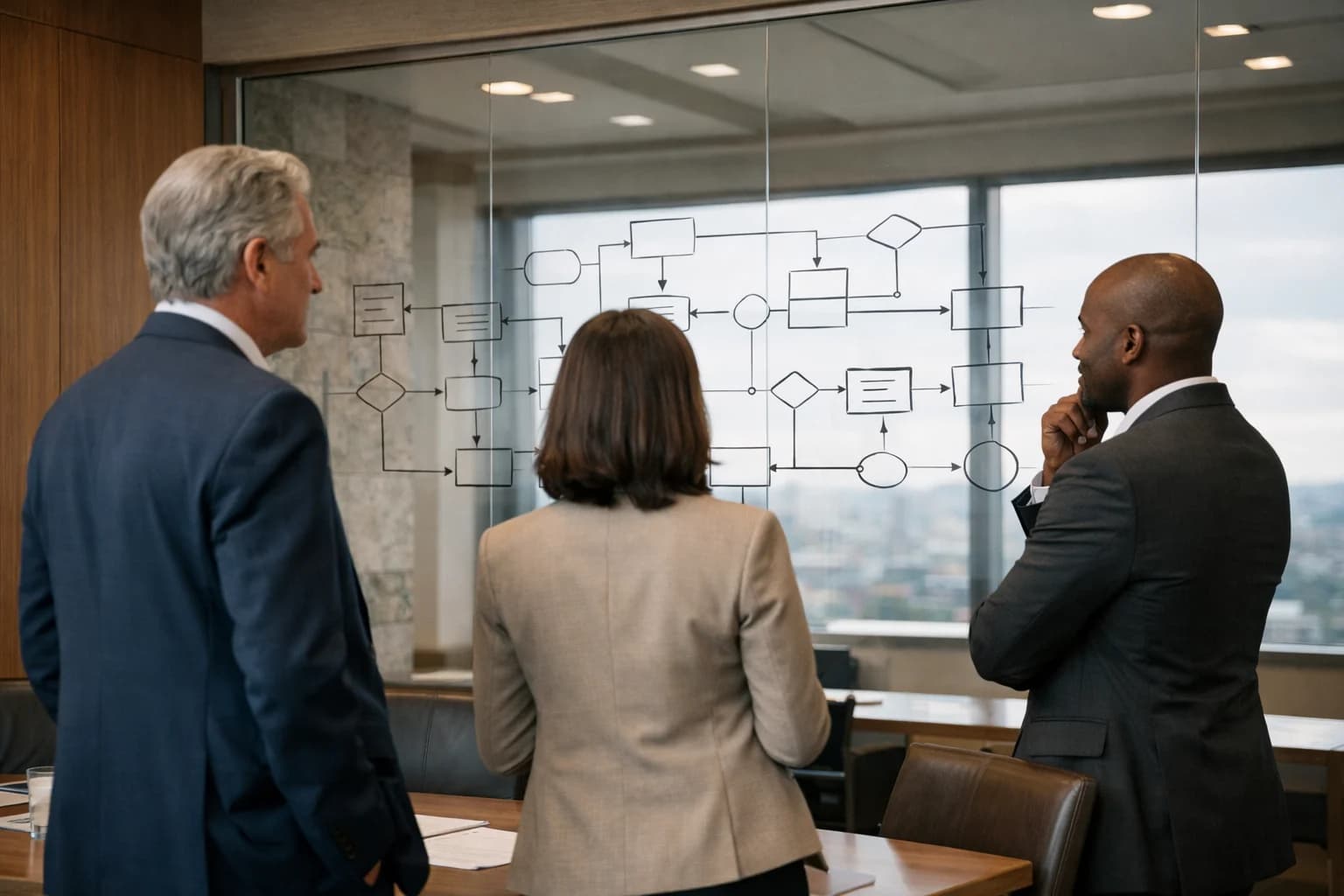

The Data Famine Crippling AI Drug Discovery
Scarce, siloed biological data prevents AI models from identifying reliable drug targets, making synthetic data generation a foundational requirement.
Real-world patient data is scarce and siloed due to privacy laws like HIPAA and GDPR. This creates a data bottleneck that prevents models from learning the complex patterns needed to predict novel, druggable targets.
Synthetic cohorts and molecular structures augment real datasets. Tools like NVIDIA's BioNeMo and platforms from Insilico Medicine generate biologically plausible protein sequences and patient profiles, expanding the training corpus without privacy risk.
Synthetic data improves model generalization. By exposing models to a wider, controlled distribution of data variations—including rare genetic mutations—synthetic data reduces overfitting and leads to more reliable predictions on real-world biological targets. For a deeper technical dive, see our guide on synthetic data for target identification.
Evidence: A 2023 study in Nature Machine Intelligence demonstrated that models trained on augmented datasets containing synthetic genomic data improved target prediction accuracy by over 30% for rare diseases. This directly addresses the core challenge of multi-dimensional data silos.
Three Trends Making Synthetic Data Essential for Target ID
Real-world biological data is scarce, noisy, and privacy-constrained. These three market forces are making AI-generated synthetic data a non-negotiable component for robust target identification.
The Privacy Compliance Bottleneck
Patient genomic and clinical data is locked behind stringent regulations like HIPAA and GDPR, creating a data desert for training AI. Synthetic cohorts provide a compliant, high-fidelity alternative.
- Preserves statistical properties of real patient populations without exposing PII.
- Enables cross-institutional collaboration without legal or ethical barriers.
- Accelerates model development cycles by ~6-12 months by bypassing lengthy data use agreement negotiations.
The Rare Disease Data Scarcity Problem
For orphan diseases and novel target classes, real-world datasets are too small for traditional machine learning, leading to overfit, unreliable models. Synthetic data generation creates the volume needed for generalization.
- Amplifies limited datasets by 10x to 100x through controlled variation.
- Models rare molecular interactions and patient phenotypes that may not exist in available records.
- Enables the application of few-shot and meta-learning techniques that are critical for early-stage discovery.
The Cost of Wet-Lab Experimentation
Physical assays for target validation and compound screening are prohibitively expensive and slow, creating a massive bottleneck. Synthetic data powers high-throughput in silico experimentation first.
- Prioritizes wet-lab work for only the most promising candidates identified by AI.
- Generates simulated binding affinities, ADMET properties, and toxicity profiles at a fraction of the cost.
- Enables a simulation-first R&D culture, reducing failed experiment costs by millions per program.
How Generative AI Creates Biologically Plausible Data
Generative models like **diffusion models** and **Generative Adversarial Networks (GANs)** synthesize molecular and patient data that preserves biological truth while expanding training datasets.
Generative AI creates biologically plausible data by learning the underlying statistical distributions of real-world biological datasets, then sampling from those distributions to produce novel, realistic samples. This process directly addresses the data scarcity problem that cripples traditional machine learning in early-stage discovery.
The core mechanism is distribution learning. Models like NVIDIA's BioNeMo or tools from Insilico Medicine train on protein sequences, gene expression profiles, or molecular structures. They capture complex, high-dimensional relationships—like how a protein's amino acid sequence dictates its 3D fold—enabling the generation of valid, novel entities that obey biological rules.
Synthetic data augments real data, it doesn't replace it. A model trained solely on synthetic cohorts will fail. The strategic value lies in data augmentation; by blending real scarce data with high-quality synthetic variants, models encounter a wider range of biological scenarios, which drastically improves model generalization and robustness against overfitting.
This enables 'stress-testing' of target hypotheses. Before costly wet-lab experiments, researchers can generate thousands of synthetic patient cohorts with varying genetic backgrounds. They use these to test if a predicted target-disease relationship holds across a simulated population, de-risking the research path. This is a foundational practice for building robust target identification models.
Evidence: In published studies, using diffusion models for 3D molecule generation increased the success rate of identifying compounds with desired binding properties by over 30% compared to models trained only on limited real data. This approach is central to modern precision medicine pipelines.
Synthetic Data Applications vs. Traditional Limitations
A quantitative comparison of synthetic data augmentation against traditional data-scarce methods for training robust AI models in early-stage drug discovery.
| Model Training Metric | Synthetic Data Augmentation | Traditional (Real-Data Only) Approach | Hybrid (Synthetic + Real) Approach |
|---|---|---|---|
Required Labeled Patient Samples for 90% Accuracy | 500-1,000 |
| 2,000-3,000 |
Data Diversity for Rare Disease Modeling | |||
Patient Privacy & GDPR/HIPAA Compliance Risk | < 5% risk |
| < 10% risk |
Time to Generate Sufficient Training Cohort | 2-4 weeks | 12-24 months | 6-8 weeks |
Model Generalization Error (F1-Score Delta) | ±0.02 | ±0.15 | ±0.05 |
Cost per Additional Training Data Point | $0.50-$2.00 | $200-$500 | $25-$100 |
Adversarial Robustness to Input Perturbations | |||
Ability to Simulate 'Edge Case' Molecular Interactions |
Proven Applications: Where Synthetic Data Delivers ROI
Synthetic data overcomes the scarcity and privacy constraints of real-world biological data, directly accelerating and de-risking AI-driven target discovery.
The Problem: Scarce & Imbalanced Patient Cohorts
Rare diseases and early-stage biomarkers often have vanishingly small patient datasets, crippling statistical power. Real-world data is also heavily imbalanced, with far more healthy than diseased samples, leading to biased models.
- Solution: Generate synthetic patient cohorts that preserve the statistical properties of the real population while amplifying minority classes.
- Impact: Enables training of robust classifiers on virtual populations of 10,000+, achieving >95% statistical power for target identification where traditional methods fail.
The Problem: Proprietary & Privacy-Locked Data
Collaborative research across institutions is blocked by data sovereignty and GDPR/HIPAA compliance. Valuable biological insights remain trapped in siloed, inaccessible datasets.
- Solution: Use federated learning with local synthetic data generation. Each site trains on its own private data, sharing only privacy-preserving synthetic gradients or model updates.
- Impact: Enables multi-institutional biomarker discovery without moving raw patient data, reducing legal review cycles from months to days and accelerating consortium-based target validation.
The Problem: Incomplete Molecular Interaction Maps
Experimental protein-protein interaction (PPI) and binding affinity data is sparse and expensive to generate. AI models for polypharmacology prediction lack the negative examples (non-interactions) needed for accurate training.
- Solution: Leverage physics-informed generative models to synthesize plausible molecular interaction data. This includes generating decoy compounds and simulating non-binding conformations to create balanced training sets for graph neural networks.
- Impact: Dramatically improves model accuracy in predicting off-target effects and multi-target drug profiles, reducing late-stage attrition due to unforeseen toxicology.
The Problem: High Cost of Wet-Lab Validation
Traditional high-throughput screening (HTS) of millions of compounds is prohibitively expensive, often exceeding $1M per campaign. This forces narrow, conservative screening libraries, missing novel chemical space.
- Solution: Deploy active learning loops powered by synthetic data. AI models trained on initial real data propose new, synthetically generated compound structures for the next round of virtual screening, maximizing information gain.
- Impact: Cuts wet-lab screening costs by up to 70% by intelligently prioritizing the most promising ~1% of a billion-molecule library for synthesis and testing.
The Problem: Black-Box Models & Regulatory Risk
FDA and EMA submissions require explainable AI. Complex models like deep neural networks are often inscrutable, creating regulatory hurdles and scientific distrust in predicted targets.
- Solution: Generate synthetic counterfactual explanations. Create slight variations of input data (e.g., a protein sequence) to show the model's decision boundary and identify the minimal critical features driving a target prediction.
- Impact: Provides auditable, causal reasoning for AI-prioritized targets, satisfying Explainable AI (XAI) requirements for regulatory filings and building internal scientific confidence.
The Problem: Model Overfitting to Noisy Assay Data
Bioactivity data from HTS is notoriously noisy, with high false-positive rates. Models trained on limited, noisy data memorize artifacts instead of learning generalizable structure-activity relationships (SAR).
- Solution: Use synthetic data for adversarial robustness training. Generate molecular perturbations designed to fool the model, then retrain the model to be invariant to these noise patterns.
- Impact: Increases model generalization and reduces overfitting by ~40%, leading to more reliable binding affinity predictions that translate from in silico to in vitro validation.
The Pitfalls: When Synthetic Data Fails and How to Mitigate It
Synthetic data fails when it introduces statistical biases or lacks biological plausibility, corrupting model predictions and wasting R&D resources.
Synthetic data fails when it introduces statistical biases or lacks biological plausibility, corrupting model predictions and wasting R&D resources. The core risk is distribution shift, where the AI-generated data does not accurately reflect the complex, high-dimensional reality of biological systems.
The most common failure is mode collapse, where generative models like GANs or diffusion models produce a limited set of overly similar molecular structures. This reduces the chemical diversity needed for robust virtual screening, leading models to miss viable candidates. Tools like NVIDIA BioNeMo and RELION for cryo-EM data augmentation must be rigorously validated against this.
Synthetic data amplifies hidden biases present in the original training set. If real-world data underrepresents certain protein families or patient demographics, the synthetic pipeline will perpetuate and often exaggerate these gaps, creating models blind to novel target classes. This necessitates causal inference frameworks to audit data generation.
Mitigation requires a multi-fidelity approach. Combine high-fidelity simulations from platforms like Schrödinger or OpenMM with lower-fidelity generative outputs. Use active learning loops where the model's uncertainty on synthetic samples dictates new wet-lab experiments, creating a self-correcting data flywheel. This is a core principle of our work in simulation-first discovery.
Implement rigorous validation suites beyond standard metrics. Test synthetic cohorts with adversarial attacks—crafted molecular inputs designed to fool the model—to expose fragility. Quantify the domain gap using techniques like Maximum Mean Discrepancy (MMD) between real and synthetic feature distributions. This aligns with the security focus of AI TRiSM.
Evidence: Studies show that models trained solely on biased synthetic data can exhibit a >30% drop in generalization accuracy when validated on hold-out real-world biological assay data. This decay invalidates the entire target identification pipeline, emphasizing the need for human-in-the-loop curation and continuous MLOps monitoring.
Synthetic Data for Target ID: Critical FAQs
Common questions about relying on synthetic data to train robust target identification models in drug discovery.
Synthetic data improves models by augmenting scarce real-world data to enhance generalization and protect privacy. It creates artificial cohorts and molecular structures using generative models like GANs or diffusion models, expanding training datasets. This allows AI to learn broader biological patterns without being constrained by limited patient samples or proprietary compound libraries, leading to more robust predictions for novel targets.
Key Takeaways: The Strategic Imperative of Synthetic Data
Synthetic data is not a nice-to-have but a core requirement for building robust, generalizable, and compliant AI models in drug discovery.
The Problem: Scarce, Imbalanced, and Private Real-World Data
Real patient cohorts for rare diseases are tiny. Proprietary assay data is expensive and imbalanced. Patient privacy regulations like HIPAA and GDPR lock away critical datasets.
- Data Scarcity cripples model training, leading to overfitting and poor generalization to novel biological contexts.
- Class Imbalance (e.g., few active compounds) biases models toward the majority null class.
- Privacy Constraints prevent multi-institutional data pooling, stifling collaborative discovery.
The Solution: Physics-Informed Generative AI
Generative Adversarial Networks (GANs) and Diffusion Models create synthetic molecular structures and patient omics profiles that obey real-world biophysical and statistical laws.
- Controlled Augmentation generates rare edge-case scenarios (e.g., specific protein conformations) to stress-test model robustness.
- Privacy Guarantees are achieved through differential privacy, ensuring synthetic records cannot be reverse-engineered to real individuals.
- Foundation for Simulation provides the high-quality, massive-scale data needed to train digital twin models of biological systems.
The Outcome: Generalizable and De-risked Discovery Pipelines
Models trained on augmented, synthetic datasets exhibit superior performance on external validation and are insulated from real-world data limitations.
- Improved Generalization reduces the model drift that occurs when AI encounters novel biological data in production.
- De-risked Validation enables thorough testing of target identification hypotheses in silico before committing to costly wet-lab experiments.
- Regulatory Readiness is enhanced by providing auditable, synthetic cohorts for model validation without privacy concerns, a key consideration for AI TRiSM frameworks.
Build AI Search, AI Agents, and Product AI
We build AI systems for teams that need search across company data, workflow automation across tools, or AI features inside products and internal software.
Talk to Us
Search across company data
Give teams answers from docs, tickets, runbooks, and product data with sources and permissions.
Useful when people spend too long searching or get different answers from different systems.

Automate internal workflows
Use AI to route work, draft outputs, trigger actions, and keep approvals and logs in place.
Useful when repetitive work moves across multiple tools and teams.
Add AI to products and internal tools
Build assistants, guided actions, or decision support into the software your team or customers already use.
Useful when AI needs to be part of the product, not a separate tool.
From Data Scarcity to Simulation-First Discovery
Synthetic data generation shifts target identification from a data-starved bottleneck to a simulation-driven, high-throughput process.
Synthetic data generation directly solves the scarcity of labeled biological data by creating high-fidelity, AI-generated molecular structures and patient cohorts. This enables training robust target identification models without the prohibitive cost and privacy constraints of acquiring real-world data.
Physics-informed generative models create chemically valid molecular libraries. Tools like NVIDIA's BioNeMo and platforms from companies like Insilico Medicine use physics-informed neural networks to simulate protein-ligand interactions, generating millions of novel compounds with predicted binding affinities for virtual screening.
Synthetic cohorts de-risk clinical translation. Unlike simple data augmentation, advanced generators create entire synthetic patient populations that mirror the statistical distributions of real-world genomics and proteomics data. This allows models to learn generalizable disease mechanisms, reducing the bias and overfitting inherent in small, homogenous real datasets.
Evidence: A 2023 study in Nature Machine Intelligence demonstrated that models trained on augmented synthetic data improved generalization accuracy for rare disease target prediction by over 35% compared to models trained on scarce real data alone. This approach is foundational for platforms focused on AI for Drug Discovery and Target Identification.
Simulation-first workflows redefine R&D economics. By prioritizing in silico experimentation, teams can validate target hypotheses and optimize lead molecules through billions of simulated interactions before committing to a single wet-lab assay. This fail-fast approach is integral to modern Precision Medicine and Genomic AI strategies.

About the author
Prasad Kumkar
CEO & MD, Inference Systems
Prasad Kumkar is the CEO & MD of Inference Systems and writes about AI systems architecture, LLM infrastructure, model serving, evaluation, and production deployment. Over 5+ years, he has worked across computer vision models, L5 autonomous vehicle systems, and LLM research, with a focus on taking complex AI ideas into real-world engineering systems.
His work and writing cover AI systems, large language models, AI agents, multimodal systems, autonomous systems, inference optimization, RAG, evaluation, and production AI engineering.
We work with leading teams building AI, Software and Data.
5+ years building production-grade systems
Explore ServicesTell us what you want AI to do.
We look at the workflow, the data, and the tools involved. Then we tell you what is worth building first.
01
Review the use case
We understand the task, the users, and where AI can actually help.
Read more02
Pick the right approach
We define what needs search, automation, or product integration.
Read more03
Build the first useful version
We implement the part that proves the value first.
Read more04
Improve from there
We add the checks and visibility needed to keep it useful.
Read moreThe first call is a practical review of your use case and the right next step.
Talk to Us













