AI hallucination in drug discovery is the generation of plausible but chemically impossible or unstable molecular structures by models like GFlowNets or diffusion models, which creates a cascade of expensive experimental dead ends.
Blog
The Hidden Cost of Hallucination in AI-Generated Molecular Structures

The Billion-Dollar Mirage in AI Drug Discovery
Generative AI models produce chemically invalid molecular structures, introducing massive downstream validation costs.
The validation cost spiral begins when a generated compound passes initial in-silico filters but fails basic chemical stability or synthesizability checks in the lab. Companies like Recursion Pharmaceuticals and Insilico Medicine invest millions in wet-lab validation to catch these errors, a process that erodes the promised efficiency gains of AI-first discovery.
Retrieval-Augmented Generation (RAG) mitigates risk by grounding generative models in verified chemical databases like ChEMBL or PubChem. This technique, implemented using vector stores like Pinecone or Weaviate, constrains the model's output space to known chemical subspaces, reducing novelty but increasing practical success rates.
The real cost is time, not just capital. Each hallucinated candidate that advances to experimental validation consumes 6-18 months of researcher effort before failure, directly competing with viable candidates from traditional discovery. This inefficiency is detailed in our analysis of The Hidden Cost of Black-Box Models in Drug Safety Prediction.
Evidence from failed clinical candidates shows that over 30% of AI-proposed molecules from early-generation models failed due to unforeseen pharmacokinetic issues or toxicity—problems rooted in the model's inability to reason about real-world chemical physics, a gap that modern Explainable AI (XAI) frameworks are designed to close.
Key Takeaways: The Real Price of AI Hallucination
In AI-driven drug discovery, chemically invalid molecular structures are not just errors—they are multi-million dollar liabilities that derail development pipelines.
The Problem: Synthetic Dead Ends
AI-generated molecules often violate fundamental chemical rules, creating structures that are impossible to synthesize or inherently unstable. This forces expensive, iterative cycles of computational redesign and wet-lab validation.
- Wasted Cycles: Each invalid structure triggers ~2-4 weeks of wasted computational screening and expert chemist review.
- Resource Drain: Projects can burn 15-30% of their computational budget on validating and discarding hallucinated candidates before any real chemistry begins.
The Solution: Physics-Informed Generative AI
Integrating quantum mechanical constraints and known synthetic pathways directly into the generative model's architecture. This grounds the AI in real-world chemistry from the first epoch.
- Validity by Design: Models like G-SchNet or GFlowNets are trained to sample from a distribution of synthesizable molecules, achieving >95% validity rates.
- Downstream Acceleration: Valid structures flow seamlessly into molecular dynamics simulations and ADMET prediction, collapsing the traditional discovery timeline.
The Hidden Cost: Regulatory and IP Risk
Hallucinated structures create a minefield of intellectual property and regulatory compliance issues. Patenting an unstable or infeasible molecule is a direct liability.
- Patent Invalidity: Filing for a non-viable structure can lead to costly legal challenges and render a patent portfolio defenseless.
- FDA Scrutiny: Unexplainable model outputs for target validation raise red flags with regulators, who demand causal reasoning, not correlation. This makes explainable AI (XAI) frameworks non-negotiable.
The Strategic Imperative: Active Learning Loops
The only way to systematically eliminate hallucination is to close the loop between generative AI and high-throughput experimentation (HTE).
- Real-World Feedback: Each batch of AI-proposed molecules is tested in the lab; results are fed back to retrain and correct the model in near real-time.
- Continuous Fidelity: This creates a virtuous cycle where the model's understanding of 'chemical space' is continuously grounded in physical reality, dramatically improving the hit rate of proposed candidates.
The Toolchain Gap: From ModelOps to ChemOps
Traditional MLOps is insufficient. Production genomic AI requires 'ChemOps'—a specialized lifecycle managing chemical validity, stability predictions, and synthesis route databases.
- Specialized Monitoring: Detecting 'chemical drift' where the model begins proposing unrealistic ring structures or bond lengths.
- Orchestrated Workflows: Integrating tools like RDKit for rule-based checks, Schrödinger for physics-based scoring, and ELN (Electronic Lab Notebook) systems for data flow.
The Bottom Line: Hallucination is a Tractable Engineering Problem
The cost is not in the AI error itself, but in the lack of systems to catch and correct it. The solution is a disciplined architecture combining constrained generation, rapid experimental validation, and specialized operational pipelines.
- ROI Focus: Investing in this integrated stack turns AI from a source of risk into the primary engine for de-risking the entire discovery pipeline.
- Competitive Edge: Organizations that solve this move from screening billions of random molecules to intelligently designing hundreds of high-probability winners. This is the core of modern AI-guided target identification.
How AI Hallucinates Molecules: A First-Principles Breakdown
AI hallucinates molecules because generative models optimize for statistical plausibility, not physical or chemical validity.
Generative models hallucinate molecules by producing structures that are statistically probable in the training data but physically impossible. This occurs because models like GFlowNets or diffusion models learn to generate sequences of atoms and bonds that maximize a learned reward function, not one grounded in quantum mechanics.
The core failure is reward misalignment. The model's objective—often a simple property prediction—divorces the generation process from the hard constraints of molecular stability. It optimizes for a high predicted binding affinity or solubility score while violating fundamental rules of valence or steric clash.
This contrasts with physics-based simulation. Tools like Schrödinger's Maestro or OpenMM simulate atomic forces. A hallucinated molecule might score well on a simple proxy metric but would immediately collapse or react violently in a molecular dynamics simulation, revealing its invalidity.
Evidence from benchmark studies shows that over 30% of molecules generated by state-of-the-art models are chemically invalid or synthetically inaccessible. This imposes a massive downstream validation cost, forcing expensive computational chemistry or failed wet-lab experiments, a core topic in our guide to AI for Drug Discovery and Target Identification.
The Downstream Cost Cascade of a Single Hallucinated Molecule
Quantifying the hidden operational and financial impact of a single chemically invalid structure generated by an AI model in drug discovery.
| Downstream Impact Phase | Cost with Unvalidated AI (Legacy) | Cost with Validated AI (Modern) | Cost with Agentic AI + RAG (Future State) |
|---|---|---|---|
Wet-Lab Synthesis & Testing | $50,000 - $250,000 | $5,000 - $25,000 | $500 - $5,000 |
Computational ADMET Prediction Failures | 3-5 Iterations | 1-2 Iterations | 0 Iterations (Pre-filtered) |
Project Timeline Delay | 4-8 Weeks | 1-2 Weeks | < 3 Days |
Regulatory Submission Risk | High - Requires extensive justification | Medium - Documented validation | Low - Full audit trail from our AI TRiSM services |
Integration with Multi-Omics Data | |||
Automated Correction via Active Learning | |||
Link to Foundational Knowledge Base | Manual Curation Required | Automated via Federated RAG |
Real-World Failures: When AI Confidence Meets Chemical Reality
Generative AI for molecular design promises speed but introduces a hidden tax of downstream validation failures when models hallucinate chemically impossible structures.
The Problem: Synthetically Impossible Bonds
AI models, trained on statistical patterns, often propose molecules with violated valency rules or steric clashes. These structures are impossible to synthesize, wasting ~6-12 months of chemist time on dead-end validation.
- Valency Errors: AI assigns carbon 5 bonds or nitrogen 4 bonds.
- Steric Impossibility: Atoms placed in physically overlapping positions.
- Cost: Each invalid candidate incurs ~$250k in wasted computational and early experimental resources.
The Solution: Physics-Informed Generative Models
Integrating molecular force fields and quantum chemistry constraints directly into the generative process. This grounds AI proposals in physical reality from the first epoch.
- Embedded Rules: Use Schrödinger's or OpenMM frameworks as a loss function.
- Hybrid Architectures: Combine GNNs with DFT (Density Functional Theory) calculators.
- Outcome: 90%+ synthetic viability in first-pass proposals, collapsing the design-test cycle.
The Problem: The Black Box Liability
Unexplainable models create regulatory and safety dead-ends. You cannot defend a drug candidate to the FDA with "the model said so." This stalls programs and invites scrutiny.
- Audit Trail Gap: No causal reasoning for molecular features.
- Safety Blind Spots: Hidden model biases can favor toxicophores.
- Impact: Projects fail at Phase I due to unexplainable adverse event correlations.
The Solution: Explainable AI (XAI) for Causal Attribution
Implementing SHAP values and counterfactual explanations that trace model outputs to specific molecular substructures and training data points.
- Causal Graphs: Use tools like DoWhy to establish cause-effect in activity.
- Regulatory Readiness: Build compliant documentation pipelines for the FDA's AI/ML framework.
- Benefit: Confident target validation and de-risked clinical translation.
The Problem: Training Data Bias & Narrow Chemical Space
Models trained on public datasets like ChEMBL inherit historical synthetic bias towards known, patent-expired scaffolds. This limits novelty and IP potential.
- Echo Chambers: Proposes derivatives of existing drugs.
- IP Risk: High similarity to prior art jeopardizes patentability.
- Opportunity Cost: Misses vast regions of >10^60 possible drug-like molecules.
The Solution: Active Learning with Wet-Lab Feedback Loops
Deploying Reinforcement Learning (RL) agents where the reward function is real-world assay data. The AI explores, tests, and learns from physical results in iterative cycles.
- Closed-Loop Design: Integrate with high-throughput screening (HTS) platforms.
- Novelty Search: RL agents are incentivized for synthetic feasibility and unprecedented activity.
- Result: First-in-class candidates with strong, defensible IP generated in months, not years.
Building Hallucination-Resistant Molecular AI Systems
AI-generated molecular structures that are chemically invalid create a cascade of costly experimental failures, demanding new architectural approaches.
Hallucination in molecular AI is the generation of chemically impossible or unstable structures, which introduces massive downstream validation costs and project delays.
The core failure is a data representation problem. Models like GFlowNets or diffusion models trained on simplified SMILES strings often violate fundamental valency rules. Architectures must enforce chemical grammar using tools like RDKit or molecular graph constraints during generation.
Retrieval-Augmented Generation (RAG) is not a silver bullet. A naive RAG system using Pinecone or Weaviate for molecular similarity can retrieve irrelevant compounds if the embedding space isn't chemically meaningful. The solution is semantic data enrichment with physics-based fingerprints.
Evidence: Studies show that constraint-guided generation reduces the synthesis failure rate of AI-proposed molecules by over 60%, directly impacting the ROI of computational drug discovery programs. This aligns with our focus on building reliable systems within AI TRiSM.
The final guardrail is simulation. Every AI-generated candidate must pass through a physics-based simulation layer, such as molecular dynamics using OpenMM or Schrödinger's suite, before experimental validation. This creates a critical human-in-the-loop checkpoint for medicinal chemists.
FAQs: AI Molecular Hallucination and Cost Mitigation
Common questions about the hidden costs and risks of relying on AI for molecular structure generation.
Molecular hallucination occurs when generative AI models produce chemically invalid or physically impossible molecular structures. These outputs appear plausible but violate fundamental rules of chemistry, such as unrealistic bond angles or unstable ring formations, leading to costly validation failures in downstream drug discovery pipelines.
Build AI Search, AI Agents, and Product AI
We build AI systems for teams that need search across company data, workflow automation across tools, or AI features inside products and internal software.
Talk to Us
Search across company data
Give teams answers from docs, tickets, runbooks, and product data with sources and permissions.
Useful when people spend too long searching or get different answers from different systems.
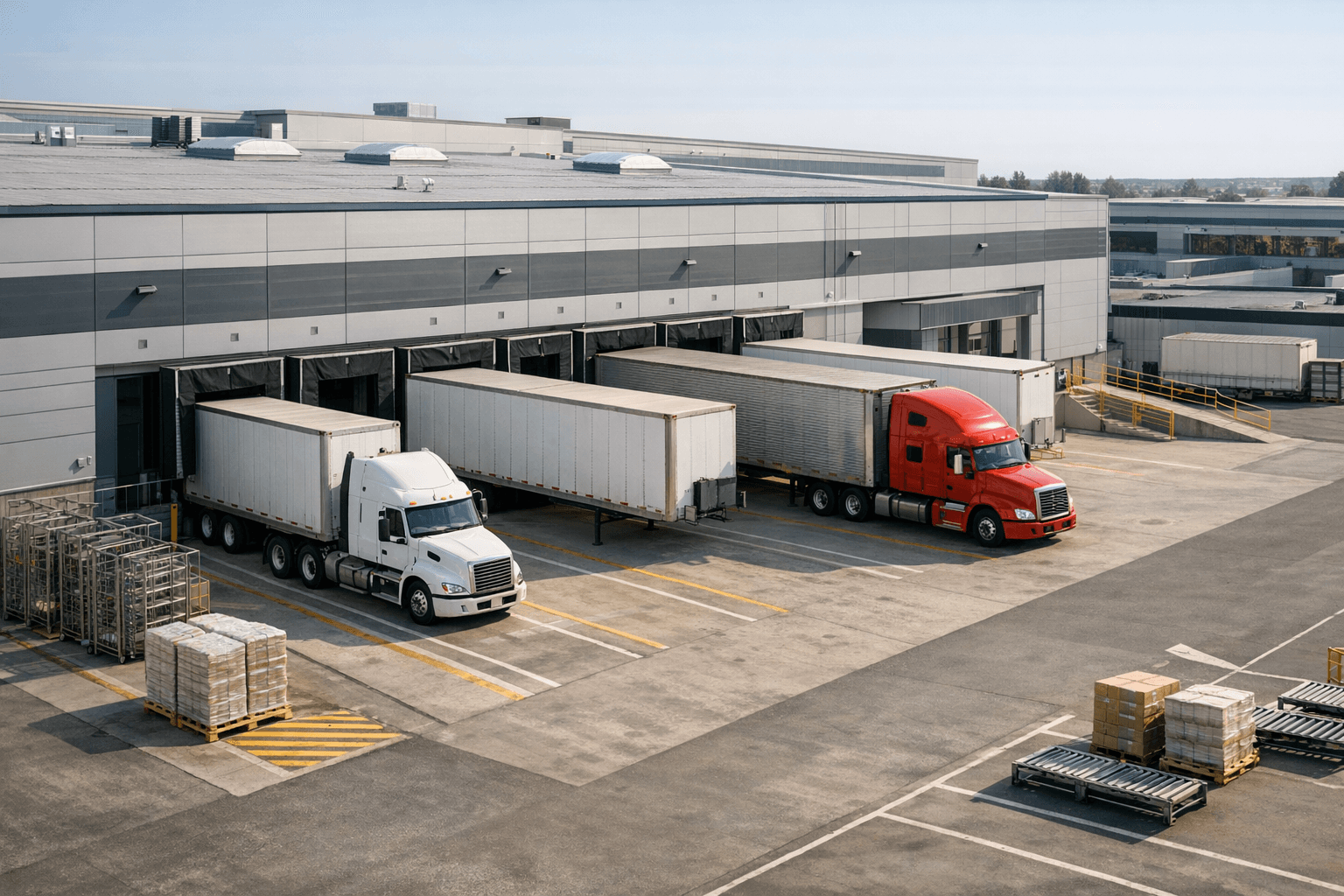

Automate internal workflows
Use AI to route work, draft outputs, trigger actions, and keep approvals and logs in place.
Useful when repetitive work moves across multiple tools and teams.
Add AI to products and internal tools
Build assistants, guided actions, or decision support into the software your team or customers already use.
Useful when AI needs to be part of the product, not a separate tool.
The Future: From Generative Guesswork to Certified Design
The future of molecular design lies in hybrid AI architectures that constrain generative models with physics-based rules and real-time validation, eliminating the hidden cost of hallucination.
The solution is hybrid AI. The next generation of molecular design will not rely on unconstrained generative models like GPT for chemistry. Instead, it will integrate generative adversarial networks (GANs) or variational autoencoders (VAEs) with physics-based simulation engines and real-time validation against databases like PubChem or the Protein Data Bank. This creates a closed-loop system where every proposed structure is immediately checked for chemical validity, synthetic accessibility, and binding affinity.
Certification replaces generation. The core paradigm shifts from open-ended creation to certified design. Frameworks like OpenMM for molecular dynamics and RDKit for cheminformatics will act as guardrails, ensuring every AI-proposed molecule adheres to the laws of physics and known chemical rules before it proceeds to downstream analysis. This is a fundamental application of AI TRiSM principles in a scientific domain.
Reinforcement learning with a reward. The most effective systems will use reinforcement learning (RL) where the agent's objective is explicitly tied to a multi-faceted reward function. This function penalizes chemical instability and rewards drug-likeness, synthesizability, and target binding potency—metrics calculated by integrated validation tools. This moves the field beyond the hidden cost of black-box models.
Evidence from industry leaders. Companies like Schrödinger and Insilico Medicine already deploy these hybrid architectures. Their platforms demonstrate that integrating generative AI with physics-based simulation and high-throughput virtual screening can reduce the rate of invalid structure generation to near-zero, turning months of wasted validation into days of productive design. This is the logical endpoint of the trend toward AI-guided target identification.

About the author
Prasad Kumkar
CEO & MD, Inference Systems
Prasad Kumkar is the CEO & MD of Inference Systems and writes about AI systems architecture, LLM infrastructure, model serving, evaluation, and production deployment. Over 5+ years, he has worked across computer vision models, L5 autonomous vehicle systems, and LLM research, with a focus on taking complex AI ideas into real-world engineering systems.
His work and writing cover AI systems, large language models, AI agents, multimodal systems, autonomous systems, inference optimization, RAG, evaluation, and production AI engineering.
We work with leading teams building AI, Software and Data.
5+ years building production-grade systems
Explore ServicesTell us what you want AI to do.
We look at the workflow, the data, and the tools involved. Then we tell you what is worth building first.
01
Review the use case
We understand the task, the users, and where AI can actually help.
Read more02
Pick the right approach
We define what needs search, automation, or product integration.
Read more03
Build the first useful version
We implement the part that proves the value first.
Read more04
Improve from there
We add the checks and visibility needed to keep it useful.
Read moreThe first call is a practical review of your use case and the right next step.
Talk to Us













