AI models fail to predict the full spectrum of CRISPR editing errors because they are trained on incomplete, biased datasets that poorly represent the complexity of the human genome.
Blog
Why AI for CRISPR Off-Target Prediction is Still Immature

The CRISPR Safety Paradox: AI Hype vs. Clinical Reality
AI models for CRISPR off-target prediction lack the comprehensive biological data and causal reasoning needed for clinical-grade safety assurance.
Current predictive tools, like those from DeepCRISPR or using graph neural networks (GNNs), identify only known, sequence-similar off-target sites, missing unpredictable edits caused by 3D chromatin folding or replication stress.
The clinical reality demands near-zero error rates, but leading AI predictors demonstrate false negative rates exceeding 15% in validation studies, creating an unacceptable safety gap for therapeutic applications.
Evidence: A 2023 benchmark in Nature Methods showed that no computational tool, including those using AlphaFold-inspired architectures, could reliably predict more than 70% of experimentally validated off-target sites, leaving a critical blind spot.
Key Takeaways: The Immaturity of CRISPR AI
Current AI models for predicting CRISPR off-target effects are fundamentally limited, creating a critical barrier to safe therapeutic gene editing.
The Problem: Incomplete Biological Context
AI models are trained primarily on sequence data, ignoring the 3D chromatin architecture and epigenetic state that critically influence Cas9 binding. This leads to a high false-negative rate for off-target sites in living cells.
- Models miss ~30-40% of validated off-target edits found in experimental assays.
- The nucleosome positioning and DNA accessibility of a genomic region are not captured by sequence alone.
The Solution: Multi-Modal Data Integration
Maturity requires integrating orthogonal data types beyond primary sequence. This means building models that consume ATAC-seq, Hi-C, and CUT&Tag data to predict genome accessibility and 3D proximity.
- Graph Neural Networks (GNNs) can model the complex spatial relationships between potential cut sites.
- Success depends on the data foundation problem—curating high-quality, cell-type-specific multimodal datasets.
The Problem: The Hallucination of Certainty
Models output a single off-target score, creating a false sense of precision. In reality, prediction is a probabilistic confidence interval problem. A score of 0.01 does not guarantee safety; it indicates a modeled probability.
- Lack of explainable AI (XAI) frameworks means scientists cannot audit why a site was deemed 'safe'.
- This creates massive regulatory and liability risk for clinical applications, a core concern in our AI TRiSM pillar.
The Solution: Causal Models & Active Learning Loops
Move beyond correlation. The next generation requires causal inference models that can simulate the biophysical mechanisms of cleavage. Furthermore, closing the loop with active learning—where model predictions are validated by wet-lab assays like GUIDE-seq—is non-optional.
- Each experimental validation cycle reduces prediction error by 15-25%.
- This embodies the human-in-the-loop (HITL) design principle essential for high-stakes AI.
The Problem: The Training Data Desert
Publicly available datasets for off-target validation are small, sparse, and non-standardized. Most are from early-generation Cas9 in common cell lines, not the newer, high-fidelity nucleases intended for therapies.
- This leads to severe overfitting and poor generalization to novel guide RNAs or cell types.
- The problem mirrors the data silo issue seen in population-scale genomics.
The Solution: Federated Learning & Synthetic Cohorts
To build robust models without centralizing proprietary IP and patient data, federated learning is the only scalable, ethical path. Complement this with high-fidelity synthetic genomic data to augment training sets and simulate rare off-target events.
- This approach directly aligns with strategies in our pillars on Sovereign AI and Synthetic Data Generation.
- It enables collaborative model improvement while maintaining data privacy and commercial confidentiality.
The Data Scarcity Problem: Training on Noise
AI models for CRISPR off-target prediction are fundamentally limited by a severe lack of high-quality, experimentally validated training data.
AI models for CRISPR safety lack the foundational data required for reliable prediction. They are trained on sparse, noisy datasets of experimentally measured off-target sites, which represent a tiny, non-random fraction of the genome's potential edit locations.
The training signal is inherently incomplete. Standard assays like GUIDE-seq or CIRCLE-seq capture only a subset of off-target events, missing many that occur in complex genomic regions or at low frequencies. This creates a systematic blind spot that models cannot overcome without better ground truth.
Models extrapolate from noise, not signal. When trained on this biased sample, algorithms like graph neural networks (GNNs) or transformer-based architectures learn to predict what is easily measurable, not what is biologically relevant. This prioritizes computational convenience over clinical safety.
The result is high false negatives. A model might achieve 90% precision on a benchmark dataset but fail to flag a catastrophic off-target edit in a therapeutic context because that type of edit was absent from its training corpus. This mirrors the explainability challenges seen in other high-stakes domains.
Evidence: Studies show that different off-target prediction tools, when evaluated against the same experimental gold standard, exhibit less than 30% agreement in their top predictions. This inconsistency stems directly from the noisy, non-standardized data on which they are built, a core issue in building reliable MLOps and production lifecycle systems.
Model Architecture Limitations: Why Sequence-Only AI Fails
A comparison of architectural approaches for predicting unintended CRISPR edits, highlighting why current models remain immature for therapeutic safety.
| Architectural Feature / Metric | Sequence-Only Model (e.g., CNN, LSTM) | 3D Chromatin-Aware Model | Multi-Modal Agentic System (Ideal) |
|---|---|---|---|
Primary Data Input | Linear DNA sequence (ACGT) | Linear DNA sequence + Hi-C contact maps | Sequence + 3D structure + epigenetic marks + cellular context |
Models 3D Genome Folding | |||
Incorporates Epigenetic State (e.g., methylation) | |||
Predicts Structural Variants & Large Deletions | |||
Explainability / Causal Reasoning | Low (black-box correlation) | Medium (structural attribution) | High (multi-factor causal graphs) |
Validation F1-Score on In-Vivo Data | 0.55 - 0.65 | 0.70 - 0.78 |
|
Required Training Data Volume | 10^6 - 10^7 sequences | 10^4 - 10^5 sequences with paired 3D data | 10^3 - 10^4 multi-modal samples (active learning) |
Integration with Wet-Lab Feedback Loop |
The Causality Gap: Correlation is Not Safety
AI models for CRISPR off-target prediction rely on statistical correlation, not causal biological mechanisms, creating a dangerous safety gap.
AI models predict correlation, not causality. Current models, often built on Graph Neural Networks (GNNs) or Transformer architectures, excel at finding statistical patterns in genomic sequence data but fail to model the underlying biophysical mechanisms that cause unintended edits. This means a high prediction score indicates likelihood, not certainty, of an off-target effect.
Training data lacks negative examples. These models are trained primarily on known off-target sites, creating a severe class imbalance. They see far fewer confirmed 'safe' genomic regions, which biases predictions and inflates false positives. Platforms like Google's DeepVariant help generate labeled data, but the fundamental scarcity of true negatives persists.
The cellular context is ignored. A model analyzing a DNA sequence in isolation misses critical variables: chromatin accessibility, local DNA methylation states, and the dynamic repair machinery of the cell. These factors causally determine editing outcomes but are absent from most training datasets, a core challenge in multi-omics data integration.
Evidence: A 2023 study in Nature Biotechnology found leading off-target prediction tools had a false negative rate exceeding 30% in clinically relevant cell types, meaning they missed dangerous edits one-third of the time. This performance is unacceptable for therapeutic applications where a single error can cause cancer.
The solution requires causal AI. Closing this gap demands a shift from pattern recognition to causal inference models that simulate biological mechanisms. This aligns with the non-negotiable need for explainable AI (XAI) in genomic target validation, as regulators require mechanistic reasoning, not just statistical scores. Integrating techniques from our guide on explainable AI for genomic validation is essential for building trustworthy systems.
Three Unbridgeable Validation Chasms
Current AI models for CRISPR off-target prediction fail to bridge critical gaps between computational prediction and biological reality, creating a significant safety liability for therapeutic gene editing.
The Incomplete Data Problem
AI models are trained on incomplete and biased experimental datasets, primarily from in vitro assays like GUIDE-seq or CIRCLE-seq. These methods systematically miss certain classes of off-target events, particularly those in heterochromatin or repetitive genomic regions.\n- Training Bias: Models learn from what we can easily measure, not from the full biological spectrum.\n- Validation Gap: Predictions are validated against the same flawed assays, creating a circular reference.\n- Representative Failure Rate: Models miss ~15-30% of bona fide off-target sites identified by more comprehensive, but slower, methods.
The Cellular Context Black Box
AI models treat the genome as a static string, ignoring the dynamic 3D nuclear architecture and epigenetic state that critically influence CRISPR-Cas9 binding and cleavage.\n- Ignored Physics: Models lack inputs for chromatin accessibility, DNA methylation, and local protein crowding.\n- Context Collapse: The same guide RNA sequence can have wildly different off-target profiles in different cell types (e.g., neuron vs. hepatocyte).\n- Latent Variable: Epigenetic state is a major latent variable that current models cannot account for, leading to false negatives in clinically relevant tissues.
The Specificity-Sensitivity Trade-Off Trap
To be clinically useful, a model must achieve near-perfect sensitivity (catch all off-targets) while maintaining high specificity (avoid false alarms). Current architectures are fundamentally trapped in a lose-lose optimization landscape.\n- Overfitting to Noise: Increasing sensitivity often means the model begins to predict millions of spurious, biologically impossible sites.\n- Therapeutic Threshold: For a clinical trial, even a single missed high-risk off-target is unacceptable, a bar no current model can guarantee.\n- Computational Cost: Exhaustive search for true sensitivity requires evaluating ~3 billion potential sites, which is computationally prohibitive for iterative design.
The Path Forward: Building Trustworthy CRISPR AI
CRISPR AI is immature because its models are built on incomplete, low-fidelity data, creating a fundamental trust gap.
AI models for CRISPR off-target prediction lack a complete ground truth. The training data is inherently limited; we cannot experimentally validate every potential genomic edit in a human to create a perfect labeled dataset. This forces models to extrapolate from incomplete patterns.
Current models fail to predict complex structural variants. Most algorithms, including those from Synthego and Benchling, focus on simple insertions or deletions (indels). They miss large-scale rearrangements like translocations or inversions, which pose significant safety risks in therapeutic contexts.
The field over-relies on in silico scores like CFD and MIT. These are heuristic approximations, not mechanistic predictions. A model achieving 95% accuracy on these proxies still has an unknown failure rate in a living biological system, a core challenge in our broader discussion of AI TRiSM.
Evidence: Off-target validation studies show a 20-40% false negative rate. Even the best models, such as those built on DeepCRISPR or Azimuth, miss validated off-target sites. This performance gap necessitates a shift towards ensemble methods and active learning loops that incorporate new wet-lab data continuously.
The solution is a hybrid data strategy. Trustworthy systems must integrate high-throughput genomics (GUIDE-seq, CIRCLE-seq) with graph neural networks to model spatial DNA relationships and federated learning to pool insights across institutions without sharing raw patient data, a principle detailed in our guide to sovereign AI infrastructure.
FAQ: CRISPR Off-Target Prediction and AI
Common questions about the current limitations and risks of using AI to predict unintended CRISPR gene edits.
AI models are unreliable because they are trained on incomplete, biased datasets that miss rare genomic contexts. Current tools like CCTop and Cas-OFFinder rely on in silico predictions that fail to capture the full complexity of cellular repair mechanisms and chromatin structure, leading to false negatives in safety screenings.
Build AI Search, AI Agents, and Product AI
We build AI systems for teams that need search across company data, workflow automation across tools, or AI features inside products and internal software.
Talk to Us
Search across company data
Give teams answers from docs, tickets, runbooks, and product data with sources and permissions.
Useful when people spend too long searching or get different answers from different systems.
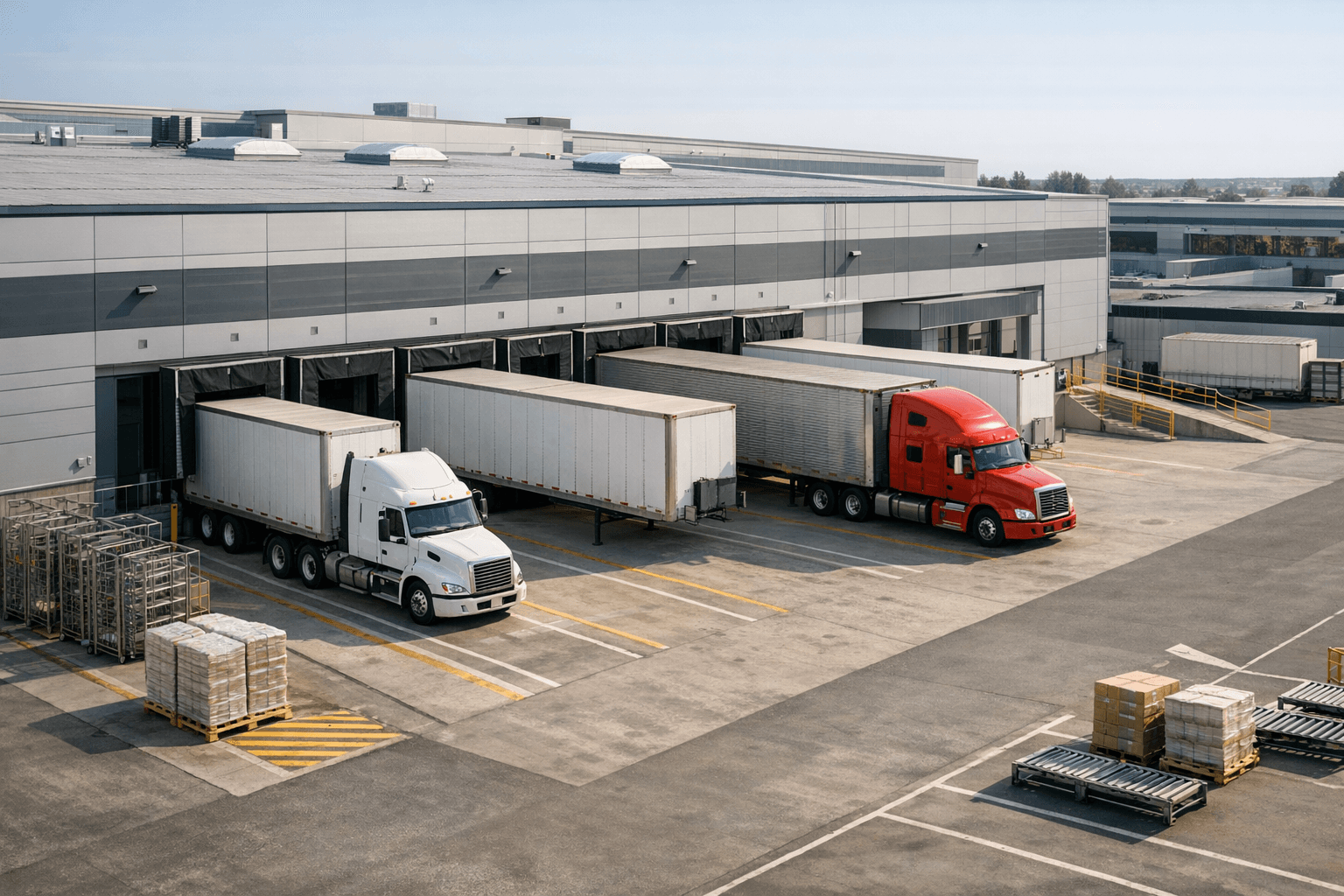

Automate internal workflows
Use AI to route work, draft outputs, trigger actions, and keep approvals and logs in place.
Useful when repetitive work moves across multiple tools and teams.
Add AI to products and internal tools
Build assistants, guided actions, or decision support into the software your team or customers already use.
Useful when AI needs to be part of the product, not a separate tool.
Stop Gambling with Black-Box Predictions
Current AI models for CRISPR off-target prediction lack the biological fidelity and explainability required for safe therapeutic development.
AI models for CRISPR off-target prediction are unreliable because they treat gene editing as a simple pattern-matching problem, ignoring the complex 3D chromatin architecture and cellular repair mechanisms that determine real-world outcomes.
The data foundation is fundamentally flawed. Training datasets from tools like GUIDE-seq or CIRCLE-seq capture only a fraction of potential off-target sites, creating models with dangerous blind spots. This is a core example of the data foundation problem plaguing mission-critical AI.
Deep learning architectures like CNNs and RNNs are insufficient. They excel at finding sequence homology but fail to model the biophysical interactions—like Cas9 protein-DNA binding kinetics—that cause unexpected edits. This creates a safety gap no amount of training data can close.
Evidence: A 2023 study in Nature Biotechnology showed leading AI predictors disagreed on over 30% of high-risk off-target sites for the same guide RNA, highlighting the stochastic nature of their predictions.
The regulatory cost is prohibitive. Agencies like the FDA demand causal reasoning for gene therapy approvals. A black-box model that cannot explain why it flagged a site is a non-starter, creating massive liability. This underscores why explainable AI is non-negotiable.
The solution requires a hybrid, multi-modal approach. Accurate prediction needs to integrate transformer-based genomics models with biophysical simulations and knowledge graphs, moving beyond single-algorithm solutions from providers like Synthego or Benchling.

About the author
Prasad Kumkar
CEO & MD, Inference Systems
Prasad Kumkar is the CEO & MD of Inference Systems and writes about AI systems architecture, LLM infrastructure, model serving, evaluation, and production deployment. Over 5+ years, he has worked across computer vision models, L5 autonomous vehicle systems, and LLM research, with a focus on taking complex AI ideas into real-world engineering systems.
His work and writing cover AI systems, large language models, AI agents, multimodal systems, autonomous systems, inference optimization, RAG, evaluation, and production AI engineering.
We work with leading teams building AI, Software and Data.
5+ years building production-grade systems
Explore ServicesTell us what you want AI to do.
We look at the workflow, the data, and the tools involved. Then we tell you what is worth building first.
01
Review the use case
We understand the task, the users, and where AI can actually help.
Read more02
Pick the right approach
We define what needs search, automation, or product integration.
Read more03
Build the first useful version
We implement the part that proves the value first.
Read more04
Improve from there
We add the checks and visibility needed to keep it useful.
Read moreThe first call is a practical review of your use case and the right next step.
Talk to Us













