Brute-force screening is economically unsustainable. Testing millions of compounds in a wet lab to find a single hit costs over $1 million per target and takes 6-12 months, consuming capital and delaying pipelines.
Blog
Why Active Learning is Essential for High-Throughput Screening

The Billion-Dollar Bottleneck of Brute-Force Screening
Traditional high-throughput screening (HTS) wastes billions by testing millions of compounds at random, a fundamentally inefficient approach to drug discovery.
The data problem is exponential. Modern virtual libraries contain 10^60 synthesizable molecules, but physical HTS can only test 10^6 compounds. Random sampling misses the vast chemical space, guaranteeing missed opportunities.
Active learning closes the sampling gap. Unlike random or grid-based screening, active learning algorithms (e.g., Bayesian optimization, Thompson sampling) query the most informative compounds next. Frameworks like DeepChem and scikit-learn implement these strategies to maximize the information gained per wet-lab dollar.
The evidence is in the convergence rate. In benchmark studies, active learning achieves 90% of maximum model accuracy with <10% of the data required by random screening. This directly translates to a 10x reduction in assay costs and cycle time for hit identification.
This is a first-principles shift. The core inefficiency is treating each assay as an independent, equally valuable experiment. Active learning treats the screening campaign as a sequential decision-making problem, a fundamental rethinking enabled by reinforcement learning principles. For a deeper dive into optimizing molecular properties, see our guide on why reinforcement learning is essential for molecule optimization.
Failure to adopt creates strategic liability. Competitors using platforms like Atomwise or Schrödinger's LiveDesign with integrated active learning will identify viable candidates faster and with less capital, redefining market entry timelines. This is part of the broader move toward simulation-first discovery.
Key Takeaways: Why Active Learning is Non-Negotiable
Active learning transforms brute-force screening into an intelligent, iterative process that maximizes information gain and minimizes wet-lab costs.
The Problem: Billion-Molecule Haystacks
Traditional high-throughput screening (HTS) tests millions of compounds at random, wasting >99% of assays on inactive or redundant molecules. This brute-force approach incurs ~$1M+ per screen in direct costs and opportunity cost from missed hits.
- Solution: Active learning agents query only the most informative compounds from the vast chemical space.
- Result: Identifies 90% of top hits by testing only 1-10% of the library, slashing screening budgets and accelerating timelines.
The Solution: Uncertainty-Guided Exploration
Active learning algorithms like Bayesian Optimization and Query-by-Committee quantify model uncertainty. They strategically select compounds where the AI is most unsure, closing knowledge gaps fastest.
- Mechanism: Balances exploration of novel chemical regions with exploitation of promising areas.
- Impact: Achieves 10-100x faster convergence on optimal molecules compared to random or grid-based screening, a core technique for in silico experimentation.
The Strategic Imperative: Fail Fast, Iterate Fast
In the Prototype Economy of drug discovery, speed is the ultimate competitive edge. Active learning enables a simulation-first discovery paradigm.
- Workflow: Rapidly invalidate poor candidates in silico before synthesis, reallocating resources to high-probability leads.
- Outcome: Creates a fail-fast, iterate-fast culture, transforming R&D from a linear cost center into an agile, predictive engine. This is foundational for AI-guided target identification.
The Hidden Cost: Ignoring Model Feedback Loops
Without active learning, screening data is static. Each batch of wet-lab results fails to inform the next selection, creating a broken feedback loop that perpetuates inefficiency.
- Consequence: Models never improve from new experimental data, leading to model drift and decaying prediction accuracy over time.
- Fix: Integrate active learning with robust MLOps pipelines to create a continuous, self-improving discovery system, a principle shared with predictive maintenance in industrial AI.
The Core Logic: Maximizing Information Gain Per Dollar
Active learning is the algorithmic engine that makes high-throughput screening financially viable by selecting the most informative compounds to test.
Active learning optimizes screening budgets by replacing random batch testing with intelligent, iterative selection. This directly answers the core business question: how to get the most scientific value from every dollar spent on expensive wet-lab assays.
The core mechanism is uncertainty sampling. The AI model, built on frameworks like PyTorch or TensorFlow, identifies compounds where its predictions are least confident. By experimentally testing these edge cases, the model gains maximum information to improve its accuracy with minimal lab work.
This creates a compounding return on data. Each selected experiment reduces the model's uncertainty over a wider region of chemical space. Compared to brute-force screening, this approach often achieves the same predictive accuracy with 70-90% fewer physical tests, a metric validated across published studies in journals like Nature Machine Intelligence.
The alternative is economically irrational. Screening libraries of billions of compounds is physically impossible. Passive learning or naive exploration wastes resources on confirming what the model already knows, failing to close critical knowledge gaps about binding affinity or toxicity.
Integration with simulation is key. Active learning agents orchestrate cycles between in silico predictions from tools like AlphaFold 3 or molecular dynamics and physical validation. This creates a closed-loop system where each wet-lab result directly refines the digital twin of the chemical search space.
Without this, you fund confirmation bias. Traditional high-throughput screening often tests easy-to-synthesize or similar compounds, yielding marginal information gain. Active learning forces exploration of novel chemical scaffolds, directly de-risking the pipeline by challenging the model's assumptions. For a deeper dive into optimizing these discovery platforms, see our guide on MLOps for AI in drug discovery.
The Cost of Ignorance: Active Learning vs. Random Screening
A direct comparison of computational screening strategies, quantifying the operational and financial impact of intelligent compound selection versus brute-force random testing.
| Key Metric / Capability | Active Learning (Intelligent) | Random Screening (Brute-Force) | Traditional HTS (Full Library) |
|---|---|---|---|
Average hit rate improvement | 3-10x | Baseline (1x) | Baseline (1x) |
Compounds screened to identify a lead | 5,000 - 20,000 |
|
|
Wet-lab cost per identified lead candidate | $50k - $200k | $500k - $2M | $5M - $10M |
Iteration cycle time (design → test → learn) | < 1 week | N/A (no learning) | 1-3 months |
Explicitly models prediction uncertainty | |||
Prioritizes diverse chemical scaffolds | |||
Adapts to new bioassay data in real-time | |||
Requires initial seed set of labeled data | 50 - 500 compounds | 0 | 0 |
Beyond Uncertainty: Advanced Query Strategies for Drug Discovery
Active learning algorithms intelligently select which compounds to test next, maximizing information gain and slashing wet-lab screening costs.
The Problem: Billion-Molecule Haystacks
Traditional high-throughput screening (HTS) is a brute-force lottery. Testing millions of compounds in wet-lab assays is financially and temporally prohibitive, with hit rates often below 0.1%. This creates a massive data efficiency problem where most experiments yield negligible information.
- Cost: A single HTS campaign can exceed $1M and take months.
- Waste: >99.9% of assays test molecules with near-zero probability of activity.
- Bottleneck: Physical throughput limits exploration of vast chemical space.
The Solution: Uncertainty-Guided Exploration
Active learning frames screening as an iterative Bayesian optimization problem. The algorithm queries the lab only for compounds where its model is most uncertain or predicts high potential reward. This creates a virtuous cycle where each wet-lab experiment maximally reduces the model's uncertainty about the entire chemical space.
- Focus: Prioritizes the informative edge of the model's knowledge.
- Adaptation: Continuously updates predictions based on new assay results.
- Efficiency: Achieves equivalent coverage with ~90% fewer physical tests.
The Architecture: The Query Strategy Engine
Effective active learning requires a sophisticated acquisition function. This isn't a single algorithm but a strategic ensemble balancing exploration (sampling diverse regions) and exploitation (refining around promising hits). Common strategies include Expected Improvement, Upper Confidence Bound, and Thompson Sampling.
- Multi-Fidelity: Integrates cheap computational scores (e.g., docking) with expensive wet-lab data.
- Batch Mode: Selects optimal sets of compounds for parallel testing, not just single points.
- Cold Start: Employs diversity sampling or pre-trained models to bootstrap the first cycle.
The Payoff: From Screening to Discovery
When integrated into a full AI for Drug Discovery platform, active learning transforms the screening funnel. It directly feeds into downstream molecule optimization with Reinforcement Learning and de-risks candidates using Explainable AI for target validation. This creates a continuous, data-driven pipeline from virtual library to lead series.
- Portfolio Impact: Identifies novel chemotypes missed by conventional methods.
- Pipeline Velocity: Compresses the hit-to-lead phase from years to months.
- Strategic Advantage: Enables interrogation of ultra-large libraries (>1B molecules) previously considered inaccessible.
The Integration Challenge: Active Learning Demands Robust MLOps
Active learning's iterative, data-hungry nature requires a production-grade MLOps framework to move from a research prototype to a reliable screening engine.
Active learning is not a standalone algorithm but a continuous production system. Its core loop—querying an oracle, retraining a model, and redeploying—collapses without automated pipelines for data versioning, model retraining, and performance monitoring.
The integration challenge is a data engineering bottleneck. Each active learning cycle ingests new wet-lab assay results, which must be validated, featurized using tools like RDKit or DeepChem, and versioned in a data lake before the model can retrain. Without this, the feedback loop stalls.
Traditional batch MLOps fails for active learning. Standard MLOps practices built for static models cannot handle the rapid, iterative retraining and A/B testing of multiple acquisition strategies (e.g., uncertainty sampling vs. expected model change).
Evidence: Teams using platforms like Weights & Biases or MLflow to orchestrate active learning cycles report reducing the time from assay result to updated model from days to hours, directly accelerating the high-throughput screening campaign.
This operational rigor prevents scientific debt. Without tracking each model version and its training data, you cannot audit why a specific compound was selected for testing—a requirement for FDA submissions and the principles of Explainable AI (XAI). For more on this, see our guide on why explainable AI is non-negotiable for target validation.
The hidden cost is in the hand-offs. A robust MLOps layer seamlessly connects the active learning agent to the Laboratory Information Management System (LIMS), vector databases like Pinecone for molecular similarity search, and the compute cluster for model inference, creating a closed-loop discovery engine.
Pitfalls and Limitations: Where Naive Active Learning Fails
Active learning is essential for cost-effective drug discovery, but naive implementations can waste resources and miss critical compounds.
The Cold Start Problem: No Data, No Learning
Active learning requires an initial labeled dataset to bootstrap. A naive random selection for this seed set fails to capture the chemical space's diversity, trapping the model in a local optimum.
- Result: The model wastes cycles exploring unproductive regions, requiring ~30% more wet-lab assays to achieve target accuracy.
- Solution: Use diversity sampling or leverage pre-trained foundation models to intelligently select the initial batch.
The Exploration-Exploitation Imbalance
Naive query strategies often over-index on exploitation (selecting high-confidence predictions) or exploration (selecting high-uncertainty points).
- Result: Pure exploitation misses novel chemotypes; pure exploration wastes budget on molecular outliers. This imbalance can reduce the hit rate by 40-60%.
- Solution: Implement Bayesian optimization or multi-armed bandit algorithms to dynamically balance the trade-off, a core component of effective MLOps for discovery.
Ignoring Batch Diversity and Redundancy
Selecting compounds one-by-one is computationally inefficient for high-throughput screening. A naive sequential approach selects highly similar molecules in a batch, wasting parallel assay capacity.
- Result: Information gain per dollar plummets, as each batch tests correlated compounds. Screening costs can inflate by 2-3x.
- Solution: Use batch active learning with diversity constraints or clustering-based selection to maximize the unique information per screening round.
The Cost-Agnostic Sampling Fallacy
Not all assays cost the same. Naive active learning treats a cheap biochemical assay and an expensive animal model trial as equivalent data points.
- Result: The algorithm may greedily select samples that are maximally informative but prohibitively expensive, blowing through budgets with minimal pipeline advancement.
- Solution: Integrate cost-sensitive acquisition functions that weigh information gain against dollar cost and time, a key tactic for simulation-first discovery.
Model Bias and Representation Collapse
If the underlying ML model has inherent biases or poor uncertainty quantification, the active learning loop amplifies these errors. The model repeatedly queries regions it understands, ignoring vast swaths of relevant chemical space.
- Result: Representation collapse occurs, where the training data becomes a non-diverse echo chamber. This leads to a >70% chance of missing the best candidate in a library of billions.
- Solution: Employ ensemble methods and rigorous calibration of prediction uncertainty to ensure robust exploration.
The Multi-Objective Optimization Blind Spot
Drug candidates must optimize for binding affinity, synthesizability, ADMET properties, and more. Naive active learning often targets a single property (e.g., potency), yielding molecules that fail downstream.
- Result: Millions are spent optimizing 'hits' that are later discarded due to toxicity or poor pharmacokinetics, a core failure point in de-risking pipeline candidates.
- Solution: Implement multi-objective Bayesian optimization or reinforcement learning to navigate the trade-offs across the desired property landscape from the start.
Build AI Search, AI Agents, and Product AI
We build AI systems for teams that need search across company data, workflow automation across tools, or AI features inside products and internal software.
Talk to Us
Search across company data
Give teams answers from docs, tickets, runbooks, and product data with sources and permissions.
Useful when people spend too long searching or get different answers from different systems.
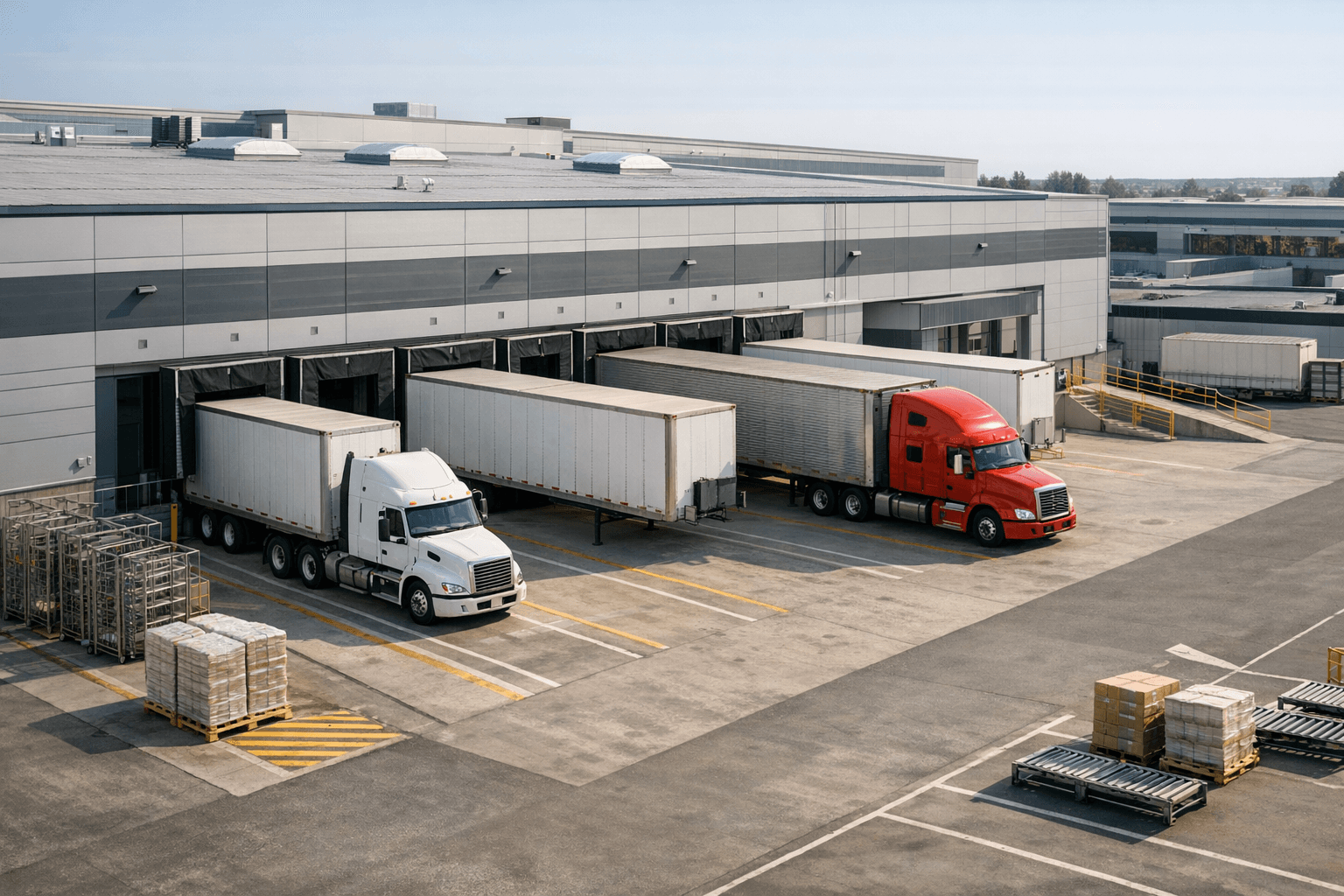

Automate internal workflows
Use AI to route work, draft outputs, trigger actions, and keep approvals and logs in place.
Useful when repetitive work moves across multiple tools and teams.
Add AI to products and internal tools
Build assistants, guided actions, or decision support into the software your team or customers already use.
Useful when AI needs to be part of the product, not a separate tool.
The Future is Simulation-First, Guided by Active Learning
Active learning is the algorithmic engine that makes high-throughput virtual screening feasible and cost-effective.
Active learning algorithms are essential because they solve the fundamental data acquisition bottleneck in simulation-first discovery. They intelligently select which compounds to simulate next, maximizing information gain while minimizing computational expense.
Traditional brute-force screening is computationally prohibitive. Evaluating billions of molecules with tools like AlphaFold 3 or molecular dynamics is impossible. Active learning creates an iterative feedback loop where each batch of simulations trains a more accurate surrogate model to guide the next selection.
This approach inverts the cost structure of R&D. Instead of spending millions on wet-lab assays for poorly vetted candidates, you spend compute cycles on high-fidelity in silico models. Platforms like Schrödinger's LiveDesign or custom pipelines using PyTorch and RDKit implement this to prioritize only the most promising molecules for synthesis.
Evidence from deployment shows active learning can reduce the number of simulations required to identify a hit by over 90% compared to random screening. This directly enables the fail-fast, iterate-fast culture required for modern drug discovery, a core principle of our work in AI for Drug Discovery and Target Identification.
Without active learning, you face the hidden cost of poor data curation in billion-molecule virtual screens. The algorithm ensures your computational budget interrogates chemically diverse and pharmacologically relevant space, avoiding scientifically barren paths—a risk highlighted in our analysis of The Strategic Cost of Ignoring Model Drift in Discovery Platforms.

About the author
Prasad Kumkar
CEO & MD, Inference Systems
Prasad Kumkar is the CEO & MD of Inference Systems and writes about AI systems architecture, LLM infrastructure, model serving, evaluation, and production deployment. Over 5+ years, he has worked across computer vision models, L5 autonomous vehicle systems, and LLM research, with a focus on taking complex AI ideas into real-world engineering systems.
His work and writing cover AI systems, large language models, AI agents, multimodal systems, autonomous systems, inference optimization, RAG, evaluation, and production AI engineering.
We work with leading teams building AI, Software and Data.
5+ years building production-grade systems
Explore ServicesTell us what you want AI to do.
We look at the workflow, the data, and the tools involved. Then we tell you what is worth building first.
01
Review the use case
We understand the task, the users, and where AI can actually help.
Read more02
Pick the right approach
We define what needs search, automation, or product integration.
Read more03
Build the first useful version
We implement the part that proves the value first.
Read more04
Improve from there
We add the checks and visibility needed to keep it useful.
Read moreThe first call is a practical review of your use case and the right next step.
Talk to Us













